
- ❓Introduction: Naive vs Graph RAG
- 🧠 Project overview
- 📊 Results
- 🔚 Conclusion
In this article, I present a comprehensive GraphRAG multi-agent system built with LangGraph that serves as an intelligent food assistant. While I’ve chosen meal planning as the demonstration domain, this architecture represents a versatile framework applicable across numerous sectors where complex, multi-faceted queries require structured knowledge retrieval.
The system handles complex scenarios across three key domains: recipe discovery with dietary constraints, shopping list generation for specific recipes, and store product location mapping within supermarkets. By combining semantic search for fuzzy matching with precise Cypher queries for structured data retrieval, the assistant performs multi-step reasoning over a Neo4j knowledge graph to deliver contextually relevant responses to nuanced queries.
1. Introduction: Naive RAG vs Graph RAG
For the purpose of this project, a Naive RAG approach is not sufficient for the following reasons:
- No Structured Relationship Modeling: Naive RAG retrieves information from unstructured text, lacking the ability to represent and reason over explicit relationships between entities. This limits its effectiveness for queries that require understanding how different pieces of information are connected.
- Limited Multi-Hop Reasoning: It processes queries at a single level, making it difficult to answer questions that require traversing multiple data points or combining information from various sources within a structured context.
- Lack of Explainability: Because retrieval is based solely on text similarity, it is challenging to trace how an answer was constructed or to provide transparent reasoning paths.
A Graph RAG system was therefore implemented to address all these issues. A graph-based framework, in fact, allows for:
- Explicit Relationship Representation: Entities and their connections are modeled directly in a knowledge graph, enabling the system to understand and utilize the structure of the data.
- Multi-Hop and Contextual Reasoning: The system can traverse the graph to perform multi-step reasoning, combining information across related nodes to answer complex queries.
- Schema-Driven Retrieval: By leveraging the graph’s schema, queries can be precisely formulated and results are consistent with the underlying data model.
- Improved Explainability: The reasoning path for each answer can be traced through the graph, providing clear explanations and greater transparency.
Such functionality makes Graph RAG systems a more robust and reliable solution for applications where structured data, complex relationships, and explainability are essential.
2. Project overview
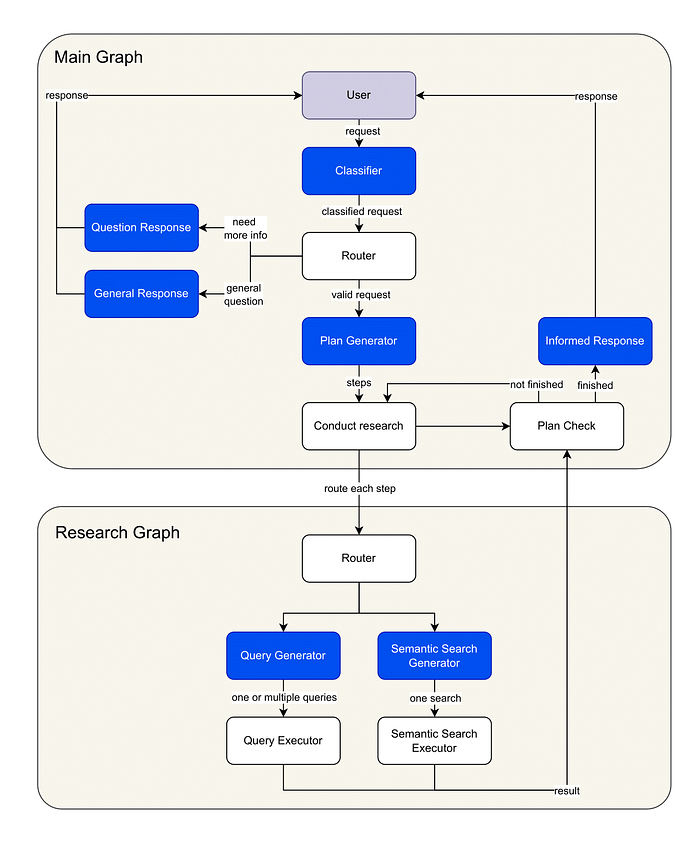
Image by author: Agentic Graph RAG diagram
GraphRAG Workflow Steps:
- Query Analysis and Routing: The user’s request is first analyzed and classified, allowing the system to route it to the appropriate workflow node. Depending on the query, the system may proceed to the next step (research plan generation), prompt the user for clarification, or respond immediately if the request is out of scope.
- Research Plan Generation: The system constructs a detailed, step-by-step research plan tailored to the complexity of the user’s query. This plan outlines the specific actions required to fulfill the request.
- Research Graph Execution: For each step in the research plan, a dedicated subgraph is invoked. The system generates Cypher queries via LLMs, targeting the Neo4j knowledge graph. Relevant nodes and relationships are retrieved using a hybrid approach that combines semantic search and structured graph queries, ensuring both breadth and precision in the results.
- Answer Generation: Leveraging the retrieved graph data, the system synthesizes a comprehensive response using an LLM, integrating information from multiple sources as needed.
For the creation of the graph, there are different choices based on your needs. I built the graph by myself with sample data to go faster but various tools can be used. In the following we see one technique using an LLM, with Langchain.
Building Neo4J Graph using an LLM
The selection of the LLM model significantly influences the output by determining the accuracy and nuance of the extracted graph data.
import os
from langchain_openai import ChatOpenAI
os.environ["OPENAI_API_KEY"] = "your-openai-api-key"
llm = ChatOpenAI(temperature=0, model_name="gpt-4o")
TheLLMGraphTransformerconverts text documents into structured graph documents by leveraging a LLM to parse and categorize entities and their relationships.
We have the flexibility to define specific types of nodes and relationships for extraction according to our requirements.
For example, we might want to have the following nodes:
RecipeFoodproduct
And the following relationships:
CONTAINS
We could specify those in this way:
from langchain_experimental.graph_transformers import LLMGraphTransformer
llm_transformer_filtered = LLMGraphTransformer(
llm=llm,
allowed_nodes=["Recipe", "Foodproduct"],
allowed_relationships=["CONTAINS"],
)
Now, we can pass in example text and examine the results.
from langchain_core.documents import Document
text = """
My favorite culinary creation is the irresistible Vegan Chocolate Cake Recipe. This delightful dessert is celebrated for its intense cocoa flavor and its incredibly soft and moist texture. It's completely vegan, dairy-free, and, thanks to the use of a special gluten-free flour blend, also gluten-free.
To make this cake, the recipe contains the following food products with their respective quantities: 250 grams of gluten-free flour blend, 80 grams of high-quality cocoa powder, 200 grams of granulated sugar, and 10 grams of baking powder. To enrich the taste and ensure a perfect rise, the recipe also contains 5 grams of vanilla extract. Among the liquid ingredients, 240 ml of almond milk and 60 ml of vegetable oil are needed.
This recipe produces a single chocolate cake, considered a FoodProduct of type dessert.
"""
documents = [Document(page_content=text)]
graph_documents_filtered = await llm_transformer_filtered.aconvert_to_graph_documents(
documents
)
print(f"Nodes:{graph_documents_filtered[0].nodes}")
print(f"Relationships:{graph_documents_filtered[0].relationships}")
And we obtain:
Nodes:[Node(id='Vegan Chocolate Cake Recipe', type='Recipe', properties={}), Node(id='Gluten-Free Flour Blend', type='Foodproduct', properties={}), Node(id='High-Quality Cocoa Powder', type='Foodproduct', properties={}), Node(id='Granulated Sugar', type='Foodproduct', properties={}), Node(id='Baking Powder', type='Foodproduct', properties={}), Node(id='Vanilla Extract', type='Foodproduct', properties={}), Node(id='Almond Milk', type='Foodproduct', properties={}), Node(id='Vegetable Oil', type='Foodproduct', properties={}), Node(id='Chocolate Cake', type='Foodproduct', properties={})]
Relationships:[Relationship(source=Node(id='Vegan Chocolate Cake Recipe', type='Recipe', properties={}), target=Node(id='Gluten-Free Flour Blend', type='Foodproduct', properties={}), type='CONTAINS', properties={}), Relationship(source=Node(id='Vegan Chocolate Cake Recipe', type='Recipe', properties={}), target=Node(id='High-Quality Cocoa Powder', type='Foodproduct', properties={}), type='CONTAINS', properties={}), Relationship(source=Node(id='Vegan Chocolate Cake Recipe', type='Recipe', properties={}), target=Node(id='Granulated Sugar', type='Foodproduct', properties={}), type='CONTAINS', properties={}), Relationship(source=Node(id='Vegan Chocolate Cake Recipe', type='Recipe', properties={}), target=Node(id='Baking Powder', type='Foodproduct', properties={}), type='CONTAINS', properties={}), Relationship(source=Node(id='Vegan Chocolate Cake Recipe', type='Recipe', properties={}), target=Node(id='Vanilla Extract', type='Foodproduct', properties={}), type='CONTAINS', properties={}), Relationship(source=Node(id='Vegan Chocolate Cake Recipe', type='Recipe', properties={}), target=Node(id='Almond Milk', type='Foodproduct', properties={}), type='CONTAINS', properties={}), Relationship(source=Node(id='Vegan Chocolate Cake Recipe', type='Recipe', properties={}), target=Node(id='Vegetable Oil', type='Foodproduct', properties={}), type='CONTAINS', properties={}), Relationship(source=Node(id='Vegan Chocolate Cake Recipe', type='Recipe', properties={}), target=Node(id='Chocolate Cake', type='Foodproduct', properties={}), type='CONTAINS', properties={})]
Finally, the generated graph documents can be stored in the Neo4J graph database, initializing it via Neo4jGraphusing the add_graph_documents method.
import os
from langchain_neo4j import Neo4jGraph
os.environ["NEO4J_URI"] = "bolt://localhost:7687"
os.environ["NEO4J_USERNAME"] = "neo4j"
os.environ["NEO4J_PASSWORD"] = "password"
graph = Neo4jGraph(refresh_schema=False)
graph.add_graph_documents(graph_documents_filtered)
Finally, we can query the graph directly from the Neo4J console to see its content:
MATCH p=(r:Recipe)-[:CONTAINS]->(fp:Foodproduct) RETURN p LIMIT 25;
MATCH p=(r:Recipe)-[:CONTAINS]->(fp:Foodproduct) RETURN p LIMIT 25;
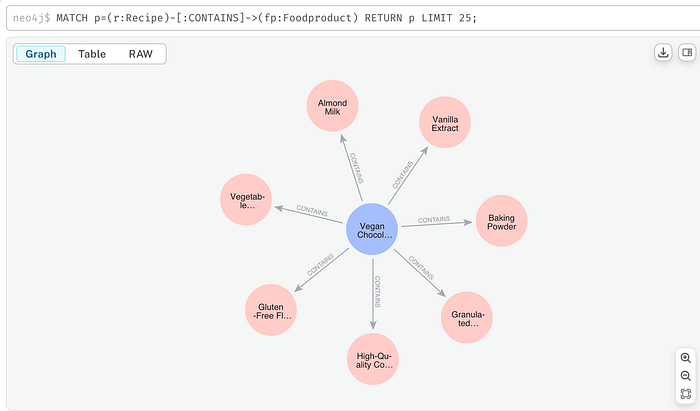

Image by author: Neo4J query response
Adding node embeddings To better understand and disambiguate user input, we can enhance our graph search with semantic search when needed. The following is just an example of how to do it, using OpenAI embeddings.
For example, if a user asks:
“Give me all the ingredients of a chocolate cake recipe that is vegan”
We need to identify the recipe node in the graph that is semantically closest to the query. To do this, we store an embedding for each Recipe node, computed from its ID.
Here’s how to generate and store an embedding in Neo4j:
import openai
from neo4j import GraphDatabase
driver = GraphDatabase.driver("bolt://localhost:7687", auth=("neo4j", "password"))
recipe_id = "Vegan Chocolate Cake Recipe"
recipe_embedding = openai.embeddings.create(model="text-embedding-3-small", input=recipe_id).data[0].embedding
with driver.session() as session:
# Create the embedding field
session.run(
"MATCH (r:Recipe {id: $recipe_id}) SET r.embedding = $embedding",
recipe_id=recipe_id,
embedding=recipe_embedding
)
# Create the vector index
session.run(
"CREATE VECTOR INDEX recipe_index IF NOT EXISTS FOR (r:Recipe) ON (r.embedding) OPTIONS {indexConfig: {`vector.dimensions`: 1536, `vector.similarity_function`: 'cosine'}}"
)
So that, later, we are able to perform semantic search:
query = "a chocolate cake recipe that is vegan"
query_embedding = openai.embeddings.create(
model="text-embedding-3-small",
input=query
).data[0].embedding
with driver.session() as session:
result = session.run(
"""
CALL db.index.vector.queryNodes('recipe_index', 1, $embedding)
YIELD node, score
RETURN node.id AS name, score
ORDER BY score DESC
""",
embedding=query_embedding
)
for record in result:
print(record["name"], "=>", record["score"])
Vegan Chocolate Cake Recipe => 0.9284169673919678
This was just a brief overview, to explore more about this technique check the Langchain documentation here, or explore other tools like the official Neo4j LLM Knowledge Graph Builder.
As I said, I created the graph iteratively introducing sample data.
Designing the workflow
The implemented system includes two graphs:
- A researcher subgraph, responsible for generating multiple Cypher queries that are used to retrieve the relevant nodes and relationships from the Neo4j knowledge graph.
- A main graph, which contains the primary workflow, including analyzing the user’s query, generating the necessary steps to accomplish the task, and producing the final response.
Main Graph structure
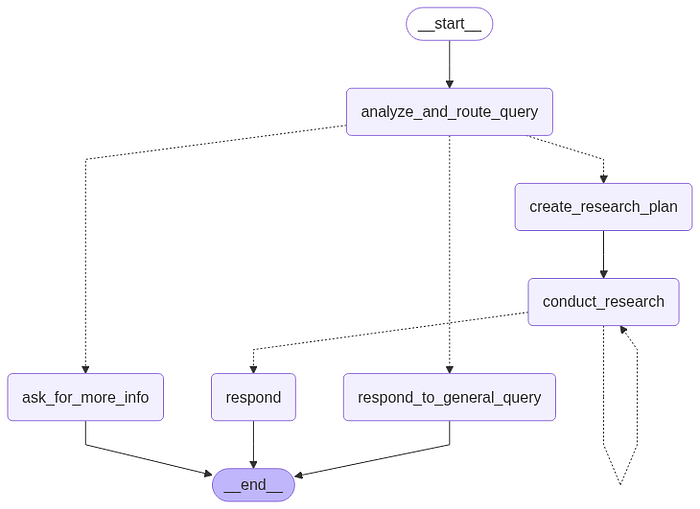
Image by author: LangGraph graph preview
One of the central concepts of LangGraph is state. Each graph execution creates a state that is passed between nodes in the graph as they execute, and each node updates this internal state with its return value after it executes.
Let’s start the project with building the graph states. To achieve this, we define the two classes:
- Router: Contains the result of classifying the user’s query into one of the classes: “more-info,” “valid,” or “general.”
from typing import Literal
from pydantic import BaseModel
class Router(BaseModel):
"""Classify user query."""
logic: str
type: Literal["more-info", "valid", "general"]
The defined graph states are:
- InputState: Includes the list of messages exchanged between the user and the agent.
- AgentState: Contains the
Router’s classification of the user’s query, the list of steps to execute in the Research Plan, the list of retrieved graph knowledge that the agent can reference.
from dataclasses import dataclass
from typing import Annotated
from langchain_core.messages import AnyMessage
from langgraph.graph import add_messages
@dataclass(kw_only=True)
class InputState:
"""
Represents the input state containing a list of messages.
Attributes:
messages (list[AnyMessage]): The list of messages associated with the state,
processed using the add_messages function.
"""
messages: Annotated[list[AnyMessage], add_messages]
from dataclasses import dataclass, field
from typing import Annotated
from utils.utils import update_knowledge
from core.state_graph.states.main_graph.input_state import InputState
from core.state_graph.states.main_graph.router import Router
from core.state_graph.states.step import Step
@dataclass(kw_only=True)
class AgentState(InputState):
"""
Represents the state of an agent within the main state graph.
Attributes:
router (Router): The routing logic for the agent.
steps (list[Step]): The sequence of steps taken by the agent.
knowledge (list[dict]): The agent's accumulated knowledge, updated via the update_knowledge function.
"""
router: Router = field(default_factory=lambda: Router(type="general", logic=""))
steps: list[Step] = field(default_factory=list)
knowledge: Annotated[list[dict], update_knowledge] = field(default_factory=list)
Step 1: Analyze and route query
The function analyze_and_route_query returns and updates the router variable of the state AgentState. The function route_query determines the next step based on the previous query classification.
Specifically, this step updates the state with a Router object whose type variable contains one of the following values: "more-info", "valid", or "general". Based on this information, the workflow will be routed to the appropriate node (one of "create_research_plan", "ask_for_more_info", or "respond_to_general_query").
async def analyze_and_route_query(state: AgentState, *, config: RunnableConfig) -> dict[str, Router]:
"""
Analyzes the current agent state and determines the routing logic for the next step.
Args:
state (AgentState): The current state of the agent, including messages and context.
config (RunnableConfig): Configuration for the runnable execution.
Returns:
dict[str, Router]: A dictionary containing the updated router object.
"""
model = init_chat_model(
name="analyze_and_route_query", **app_config["inference_model_params"]
)
messages = [{"role": "system", "content": ROUTER_SYSTEM_PROMPT}] + state.messages
print("---ANALYZE AND ROUTE QUERY---")
print(f"MESSAGES: {state.messages}")
response = cast(
Router, await model.with_structured_output(Router).ainvoke(messages)
)
return {"router": response}
def route_query(state: AgentState) -> Literal["create_research_plan", "ask_for_more_info", "respond_to_general_query"]:
"""
Determines the next action for the agent based on the router type in the current state.
Args:
state (AgentState): The current state of the agent, including the router type.
Returns:
Literal["create_research_plan", "ask_for_more_info", "respond_to_general_query"]:
The next node/action to execute in the state graph.
Raises:
ValueError: If the router type is unknown.
"""
_type = state.router.type
if _type == "valid":
return "create_research_plan"
elif _type == "more-info":
return "ask_for_more_info"
elif _type == "general":
return "respond_to_general_query"
else:
raise ValueError(f"Unknown router type {_type}")
Output example to the question:
"Suggest me some sweet recipe!"
{
"logic":"While the flavor profile 'Sweet' is provided, other mandatory constraints (dietary requirements, meal timing, recipe complexity, meal course, cooking time, and caloric content) are missing. Therefore, more information is needed before suggesting recipes.",
"type":"more-info"
}
The request is classified as more-info because it doesn’t contain all the mandatory constraints that we have inserted in the prompt.
Step 1.1 Out of scope / More informations needed
We then define the functions ask_for_more_info and respond_to_general_query, which directly generate a response for the user by making a call to the LLM: the first will be executed if the router determines that more information is needed from the user, while the second generates a response to a general query not related to our topic. In this case, it is necessary to concatenate the generated response to the list of messages, updating the messages variable in the state.
async def ask_for_more_info(state: AgentState, *, config: RunnableConfig) -> dict[str, list[BaseMessage]]:
"""
Asks the user for more information based on the current routing logic.
Args:
state (AgentState): The current state of the agent, including routing logic and messages.
config (RunnableConfig): Configuration for the runnable execution.
Returns:
dict[str, list[BaseMessage]]: A dictionary containing the new message(s) requesting more information.
"""
model = init_chat_model(
name="ask_for_more_info", **app_config["inference_model_params"]
)
system_prompt = MORE_INFO_SYSTEM_PROMPT.format(logic=state.router.logic)
messages = [{"role": "system", "content": system_prompt}] + state.messages
response = await model.ainvoke(messages)
return {"messages": [response]}
async def respond_to_general_query(state: AgentState, *, config: RunnableConfig) -> dict[str, list[BaseMessage]]:
"""
Generates a response to a general user query based on the agent's current state and routing logic.
Args:
state (AgentState): The current state of the agent, including routing logic and messages.
config (RunnableConfig): Configuration for the runnable execution.
Returns:
dict[str, list[BaseMessage]]: A dictionary containing the generated response message(s).
"""
model = init_chat_model(
name="respond_to_general_query", **app_config["inference_model_params"]
)
system_prompt = GENERAL_SYSTEM_PROMPT.format(logic=state.router.logic)
print("---RESPONSE GENERATION---")
messages = [{"role": "system", "content": system_prompt}] + state.messages
response = await model.ainvoke(messages)
return {"messages": [response]}
Output example to the question:
“What’s the weather like in Munich?”
{
"logic":"The request is about the current weather in Munich, which is not related to the Recipe, Shopping List, or Location Use Cases. It is therefore classified as a general question.",
"type":"general"
}
# ---RESPONSE GENERATION---
"I understand you’d like to know the weather in Munich, but I can only help with recipes, shopping lists for recipes, and the location of products in supermarkets."
Step 2: Create a research plan
If the query classification returns the value "valid", the user's request is in scope with the document, and the workflow will reach the create_research_plan node, whose function creates a step-by-step research plan for answering a food-related query.
review_research_planchecks and improves a research plan for quality and relevance.reduce_research_plansimplifies or condenses the plan’s steps to make it more efficient.create_research_planorchestrates the process: it generates a plan, reduces it, reviews it, and returns the final steps.
async def review_research_plan(plan: Plan) -> Plan:
"""
Reviews a research plan to ensure its quality and relevance.Args:
plan (Plan): The research plan to be reviewed.
Returns:
Plan: The reviewed and potentially modified research plan.
"""
formatted_plan = ""
for i, step in enumerate(plan["steps"]):
formatted_plan += f"{i+1}. ({step['type']}): {step['question']}\n"
model = init_chat_model(
name="create_research_plan", **app_config["inference_model_params"]
)
system_prompt = REVIEW_RESEARCH_PLAN_SYSTEM_PROMPT.format(
schema=neo4j_graph.get_structured_schema, plan=formatted_plan
)
reviewed_plan = cast(
Plan, await model.with_structured_output(Plan).ainvoke(system_prompt)
)
return reviewed_plan
async def reduce_research_plan(plan: Plan) -> Plan:
"""
Reduces a research plan by simplifying or condensing its steps.
Args:
plan (Plan): The research plan to be reduced.
Returns:
Plan: The reduced research plan.
"""
formatted_plan = ""
for i, step in enumerate(plan["steps"]):
formatted_plan += f"{i+1}. ({step['type']}): {step['question']}\n"
model = init_chat_model(
name="reduce_research_plan", **app_config["inference_model_params"]
)
system_prompt = REDUCE_RESEARCH_PLAN_SYSTEM_PROMPT.format(
schema=neo4j_graph.get_structured_schema, plan=formatted_plan
)
reduced_plan = cast(
Plan, await model.with_structured_output(Plan).ainvoke(system_prompt)
)
return reduced_plan
async def create_research_plan(
state: AgentState, *, config: RunnableConfig
) -> dict[str, list[str] | str]:
"""
Creates, reduces, and reviews a research plan based on the agent's current knowledge and messages.
Args:
state (AgentState): The current state of the agent, including knowledge and messages.
config (RunnableConfig): Configuration for the runnable execution.
Returns:
dict[str, list[str] | str]: A dictionary containing the final steps of the reviewed plan and an empty knowledge list.
"""
formatted_knowledge = "\n".join([item["content"] for item in state.knowledge])
model = init_chat_model(
name="create_research_plan", **app_config["inference_model_params"]
)
system_prompt = RESEARCH_PLAN_SYSTEM_PROMPT.format(
schema=neo4j_graph.get_structured_schema, context=formatted_knowledge
)
messages = [{"role": "system", "content": system_prompt}] + state.messages
print("---PLAN GENERATION---")
# Generate plan
plan = cast(Plan, await model.with_structured_output(Plan).ainvoke(messages))
print("Plan")
for i, step in enumerate(plan["steps"]):
print(f"{i+1}. ({step['type']}): {step['question']}")
# Reduce plan
reduced_plan = cast(Plan, await reduce_research_plan(plan=plan))
print("Reduced Plan")
for i, step in enumerate(reduced_plan["steps"]):
print(f"{i+1}. ({step['type']}): {step['question']}")
# Review plan
reviewed_plan = cast(Plan, await review_research_plan(plan=reduced_plan))
print("Reviewed Plan")
for i, step in enumerate(reviewed_plan["steps"]):
print(f"{i+1}. ({step['type']}): {step['question']}")
return {"steps": reviewed_plan["steps"], "knowledge": []}
Output example to the question:
“Suggest me some recipes. I’m vegetarian and I don’t know what to cook for breakfast. It should be less than 1000 calories though. I don’t have any other preferences.”
{
"steps":
[
{"type": "semantic_search", "question": "Find recipes that fit a vegetarian diet by searching for 'Vegetarian' in the Diet node's name property."},
{"type": "semantic_search", "question": "Find recipes that are suitable for breakfast by searching for 'Breakfast' in the MealMoment node's name property."},
{"type": "query_search", "question": "Retrieve recipes that are both vegetarian and served during breakfast by finding the intersection of results from steps 1 and 2. Filter these recipes to ensure they contain ingredients with a total calorie count of less than 1000. Use the CONTAINS relationship to sum the calories from the FoodProduct nodes. Limit 50."}
]
}
In this case, the user’s request requires three steps to retrieve the information.
Step 3: Conduct research
This function takes the first step from the research plan and uses it to conduct research. For the research, the function calls the subgraph researcher_graph, which returns all the new knowledge gathered that we will explore in the next section. Finally, we update the steps variable in the state by removing the step that was just executed.
async def conduct_research(state: AgentState) -> dict[str, Any]:
"""
Executes a research step using the research graph and updates the agent's knowledge.Args:
state (AgentState): The current state of the agent, including steps and knowledge.
Returns:
dict[str, Any]: A dictionary containing the updated knowledge and remaining steps.
"""
response = await research_graph.ainvoke(
{"step": state.steps[0], "knowledge": state.knowledge}
)
knowledge = response["knowledge"]
step = state.steps[0]
print(
f"\n{len(knowledge)} pieces of knowledge retrieved in total for the step: {step}."
)
return {"knowledge": knowledge, "steps": state.steps[1:]}
Step 4: Research subgraph building
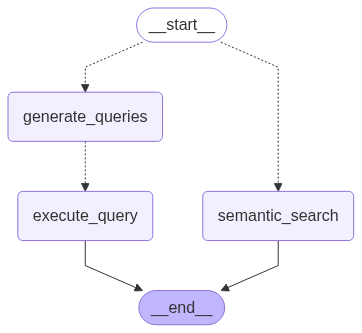
Image by author: Research Graph diagram
As visible in the image above, the graph consists of:
- a query generation and an execution step, or
- a semantic search step
As we did for the main graph, let’s proceed with defining the states QueryState (private state for the execute_query node in the researcher graph) and ResearcherState (state of the researcher graph).
@dataclass(kw_only=True)
class QueryState:
"""State class for managing research queries in the research graph."""
query: str
class Step(TypedDict):
"""Single research step"""
question: str
type: Literal["semantic_search", "query_search"]
@dataclass(kw_only=True)
class ResearcherState:
"""State of the researcher graph."""
step: Step
queries: list[str] = field(default_factory=list)
knowledge: Annotated[list[dict], update_knowledge] = field(default_factory=list)
Step 4.1 Semantic search
This step performs vector-based semantic searches on the Neo4j graph database to find relevant nodes based on similarity rather than exact matches.
It is composed by two functions:
semantic_searchdetermines search parameters using an LLM and orchestrates the semantic search execution.execute_semantic_searchperforms the actual vector similarity search using OpenAI embeddings and Neo4j’s vector index.
def execute_semantic_search(node_label: str, attribute_name: str, query: str):
"""Execute a semantic search on Neo4j vector indexes.
This function performs vector-based similarity search using OpenAI embeddings
to find nodes in the Neo4j graph database that are semantically similar to
the provided query. It converts the query to an embedding vector and searches
the corresponding vector index for the most similar nodes.
Args:
node_label (str): The label of the node type to search (e.g., 'Recipe', 'FoodProduct').
attribute_name (str): The attribute/property of the node to search within (e.g., 'name', 'description').
query (str): The search query to find semantically similar content.
Returns:
list: A list of dictionaries containing the matching nodes with their attributes,
ordered by similarity score (highest first).
"""
index_name = f"{node_label.lower()}_{attribute_name}_index"
top_k = 1
query_embedding = (
openai.embeddings.create(model=app_config["embedding_model"], input=query)
.data[0]
.embedding
)
nodes = (
f"node.name as name, node.{attribute_name} as {attribute_name}"
if attribute_name != "name"
else f"node.{attribute_name} as name"
)
response = neo4j_graph.query(
f"""
CALL db.index.vector.queryNodes('{index_name}', {top_k}, {query_embedding})
YIELD node, score
RETURN {nodes}
ORDER BY score DESC"""
)
print(
f"Semantic Search Tool invoked with parameters: node_label: '{node_label}', attribute_name: '{attribute_name}', query: '{query}'"
)
print(f"Semantic Search response: {response}")
return response
async def semantic_search(state: ResearcherState, *, config: RunnableConfig):
"""Perform semantic search to find relevant nodes in the research graph.
This function analyzes a research question to determine optimal search parameters
and executes a semantic search on the Neo4j graph database. It uses an LLM to
identify which node type and attribute should be searched, then performs vector-based
similarity search to find semantically related content that can help answer the question.
Args:
state (ResearcherState): The current researcher state containing the
research step question and accumulated knowledge.
config (RunnableConfig): Configuration for the runnable execution.
Returns:
dict[str, list]: A dictionary with a "knowledge" key containing
a list with the semantic search results formatted as knowledge items.
"""
class Response(TypedDict):
node_label: str
attribute_name: str
query: str
model = init_chat_model(
name="semantic_search", **app_config["inference_model_params"]
)
vector_indexes = neo4j_graph.query("SHOW VECTOR INDEXES YIELD name RETURN name;")
print(f"vector_indexes: {vector_indexes}")
system_prompt = SEMANTIC_SEARCH_SYSTEM_PROMPT.format(
schema=neo4j_graph.get_structured_schema,
vector_indexes=str(vector_indexes)
)
messages = [
{"role": "system", "content": system_prompt},
{"role": "human", "content": state.step["question"]},
]
response = cast(
Response, await model.with_structured_output(Response).ainvoke(messages)
)
sem_search_response = execute_semantic_search(
node_label=response["node_label"],
attribute_name=response["attribute_name"],
query=response["query"],
)
search_names = [f"'{record['name']}'" for record in sem_search_response]
joined_search_names = ", ".join(search_names)
knowledge = {
"id": new_uuid(),
"content": f"Executed Semantic Search on {response['node_label']}.{response['attribute_name']} for values similar to: '{response['query']}'\nResults: {joined_search_names}",
}
return {"knowledge": [knowledge]}
Output example to the produced steps:
[
{"type": "semantic_search", "question": "Find recipes that fit a vegetarian diet by searching for 'Vegetarian' in the Diet node's name property."},
{"type": "semantic_search", "question": "Find recipes that are suitable for breakfast by searching for 'Breakfast' in the MealMoment node's name property."}
]
# -- NEW KNOWLEDGE --
Semantic Search Tool invoked with parameters: node_label: 'Diet', attribute_name: 'name', query: 'Vegetarian'
Semantic Search response: [{'name': 'Vegetarian'}]
Semantic Search Tool invoked with parameters: node_label: 'MealMoment', attribute_name: 'name', query: 'Breakfast'
Semantic Search response: [{'name': 'Breakfast'}]
Step 4.2 Generate queries
This step generate search queries based on the question (a step in the research plan). This function uses a LLM to generate diverse Cypher queries to help answer the question. It is composed by three functions:
generate_queriesmain function that generates initial queries and applies both correction methods.correct_query_by_llmcorrects Cypher queries using a language model with schema awareness.correct_query_by_parserapplies structural corrections using a parser-based query corrector.
async def correct_query_by_llm(query: str) -> str:
"""Correct a Cypher query using a language model.
This function uses an LLM to review and correct a Cypher query based on
the Neo4j graph schema. It provides schema-aware correction to ensure
the query is properly formatted and uses valid relationships and nodes.
Args:
query (str): The Cypher query to be corrected.
Returns:
str: The corrected Cypher query.
"""
model = init_chat_model(
name="correct_query_by_llm", **app_config["inference_model_params"]
)
system_prompt = FIX_QUERY_SYSTEM_PROMPT.format(
schema=neo4j_graph.get_structured_schema
)
messages = [
{"role": "system", "content": system_prompt},
{"role": "human", "content": query},
]
response = await model.ainvoke(messages)
return response.content
def correct_query_by_parser(query: str) -> str:
"""Correct a Cypher query using a parser-based corrector.
This function uses the CypherQueryCorrector to parse and correct
Cypher queries based on the graph schema. It extracts the Cypher
query from the text and applies structural corrections.
Args:
query (str): The text containing the Cypher query to be corrected.
Returns:
str: The corrected Cypher query.
"""
corrector_schema = [
Schema(el["start"], el["type"], el["end"])
for el in neo4j_graph.get_structured_schema.get("relationships", [])
]
cypher_query_corrector = CypherQueryCorrector(corrector_schema)
extracted_query = extract_cypher(text=query)
corrected_query = cypher_query_corrector(extracted_query)
return corrected_query
async def generate_queries(
state: ResearcherState, *, config: RunnableConfig
) -> dict[str, list[str]]:
"""Generate and correct Cypher queries for a research step.
This function generates multiple Cypher queries based on a research question
and existing knowledge context. It uses an LLM to generate initial queries,
then applies both LLM-based and parser-based corrections to ensure the
queries are valid and properly formatted for the Neo4j graph database.
Args:
state (ResearcherState): The current researcher state containing the
research step question and accumulated knowledge.
config (RunnableConfig): Configuration for the runnable execution.
Returns:
dict[str, list[str]]: A dictionary with a "queries" key containing
a list of corrected Cypher queries.
"""
class Response(TypedDict):
queries: list[str]
print("---GENERATE QUERIES---")
formatted_knowledge = "\n\n".join(
[f"{i+1}. {item['content']}" for i, item in enumerate(state.knowledge)]
)
model = init_chat_model(
name="generate_queries", **app_config["inference_model_params"]
)
system_prompt = GENERATE_QUERIES_SYSTEM_PROMPT.format(
schema=neo4j_graph.get_schema, context=formatted_knowledge
)
messages = [
{"role": "system", "content": system_prompt},
{"role": "human", "content": state.step["question"]},
]
response = cast(
Response, await model.with_structured_output(Response).ainvoke(messages)
)
response["queries"] = [
await correct_query_by_llm(query=q) for q in response["queries"]
]
response["queries"] = [
correct_query_by_parser(query=q) for q in response["queries"]
]
print(f"Queries: {response['queries']}")
return {"queries": response["queries"]}
Output example to the question (after the semantic search queries execution):
“Suggest me some recipes. I’m vegetarian and I don’t know what to cook for breakfast. It should be less than 1000 calories though. I don’t have any other preferences.”
MATCH (r:Recipe)-[:FITS_DIET]->(:Diet {name: 'Vegetarian'}),
(r)-[:SERVED_DURING]->(:MealMoment {name: 'Breakfast'}),
(r)-[c:CONTAINS]->(fp:FoodProduct)
WITH r, SUM(c.grams * (fp.calories / 100.0)) AS total_calories
WHERE total_calories < 1000
RETURN r.name AS recipe_name, total_calories
LIMIT 5
And after its execution:
# -- NEW KNOWLEDGE --
╒════════════════════════════╤══════════════════╕
│recipe_name │total_calories
╞════════════════════════════╪══════════════════╡
│"Mascarpone Dessert" │945.8000000000001
├────────────────────────────┼──────────────────┤
│"Buffalo Mozzarella Salad" │668.88
├────────────────────────────┼──────────────────┤
│"Raisin and Almond Snack" │374.69999999999993
├────────────────────────────┼──────────────────┤
│"Mozzarella and Basil Salad"│528.4
└────────────────────────────┴──────────────────┘
Step 4.3 Building subgraph
def build_research_graph():
builder = StateGraph(ResearcherState)
builder.add_node(generate_queries)
builder.add_node(execute_query)
builder.add_node(semantic_search)
builder.add_conditional_edges(
START,
route_step,
{"generate_queries": "generate_queries", "semantic_search": "semantic_search"},
)
builder.add_conditional_edges(
"generate_queries",
query_in_parallel, # type: ignore
path_map=["execute_query"],
)
builder.add_edge("execute_query", END)
builder.add_edge("semantic_search", END)
return builder.compile()
research_graph = build_research_graph()
Step 5: Check finished
Using a conditional_edge, we build a loop with the end condition determined by the value returned by check_finished. This function checks that there are no more steps to process in the list of steps created by the create_research_plan node. Once all steps are completed, the flow proceeds to the respond node.
def check_finished(state: AgentState) -> Literal["respond", "conduct_research"]:
"""
Determines whether the agent should respond or conduct further research based on the steps taken.Args:
state (AgentState): The current state of the agent, including the steps performed.
Returns:
Literal["respond", "conduct_research"]:
"conduct_research" if there are steps present, otherwise "respond".
"""
if len(state.steps or []) > 0:
return "conduct_research"
else:
return "respond"
Step 6: Respond
Generates a final response to the user’s query based on the conducted research. This function formulates a comprehensive answer using the conversation history and the documents retrieved by the researcher agent.
async def respond(
state: AgentState, *, config: RunnableConfig
) -> dict[str, list[BaseMessage]]:
"""
Generates a final response to the user based on the agent's accumulated knowledge and messages.
Args:
state (AgentState): The current state of the agent, including knowledge and messages.
config (RunnableConfig): Configuration for the runnable execution.
Returns:
dict[str, list[BaseMessage]]: A dictionary containing the generated response message(s).
"""
print("--- RESPONSE GENERATION STEP ---")
model = init_chat_model(name="respond", **app_config["inference_model_params"])
formatted_knowledge = "\n\n".join([item["content"] for item in state.knowledge])
prompt = RESPONSE_SYSTEM_PROMPT.format(context=formatted_knowledge)
messages = [{"role": "system", "content": prompt}] + state.messages
response = await model.ainvoke(messages)
return {"messages": [response]}
Step 7: Building main graph
def build_main_graph():
builder = StateGraph(AgentState, input=InputState)
builder.add_node(analyze_and_route_query)
builder.add_node(ask_for_more_info)
builder.add_node(respond_to_general_query)
builder.add_node(create_research_plan)
builder.add_node(conduct_research)
builder.add_node("respond", respond)
builder.add_edge("create_research_plan", "conduct_research")
builder.add_edge(START, "analyze_and_route_query")
builder.add_conditional_edges("analyze_and_route_query", route_query)
builder.add_conditional_edges("conduct_research", check_finished)
builder.add_edge("respond", END)
return builder.compile()
3. Results
We can see its performance testing it with the following question:
“Give me the shopping list for the recipe pasta alla carbonara.”
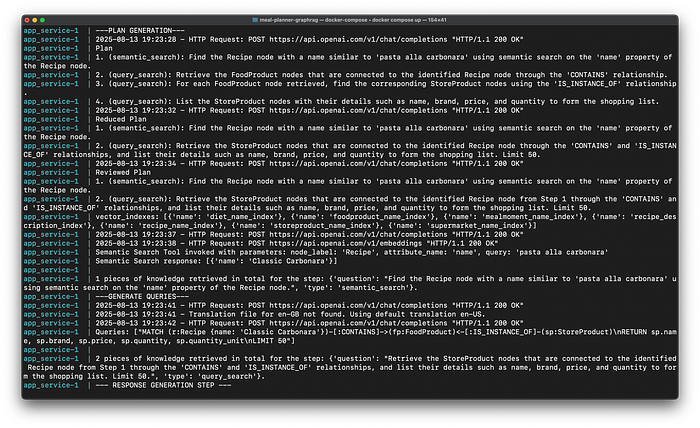
Image by author: Console Logs
As we can see from the console logs, the Main Graph create the following reviewed plan:
- semantic_search: Find the Recipe node with a name similar to ‘pasta alla carbonara’ using semantic search on the ‘name’ property of the Recipe node.
- query_search: Retrieve the StoreProduct nodes that are connected to the identified Recipe node from Step 1 through the ‘CONTAINS’ and ‘IS_INSTANCE_OF’ relationships, and list their details such as name, brand, price, and quantity to form the shopping list. Limit 50.
After executing the first step, we get to know that the exact name of the Recipe node corresponding to the ‘pasta alla carbonara’ is ‘Classic Carbonara’.
app_service-1 | Semantic Search Tool invoked with parameters: node_label: 'Recipe', attribute_name: 'name', query: 'pasta alla carbonara'
app_service-1 | Semantic Search response: [{'name': 'Classic Carbonara'}]
Then it executes the second step, with the following Cypher query:
MATCH (r:Recipe {name: 'Classic Carbonara'})-[:CONTAINS]->(fp:FoodProduct)<-[:IS_INSTANCE_OF]-(sp:StoreProduct)
RETURN sp.name, sp.brand, sp.price, sp.quantity, sp.quantity_unit
LIMIT 50
And then we get the final response.
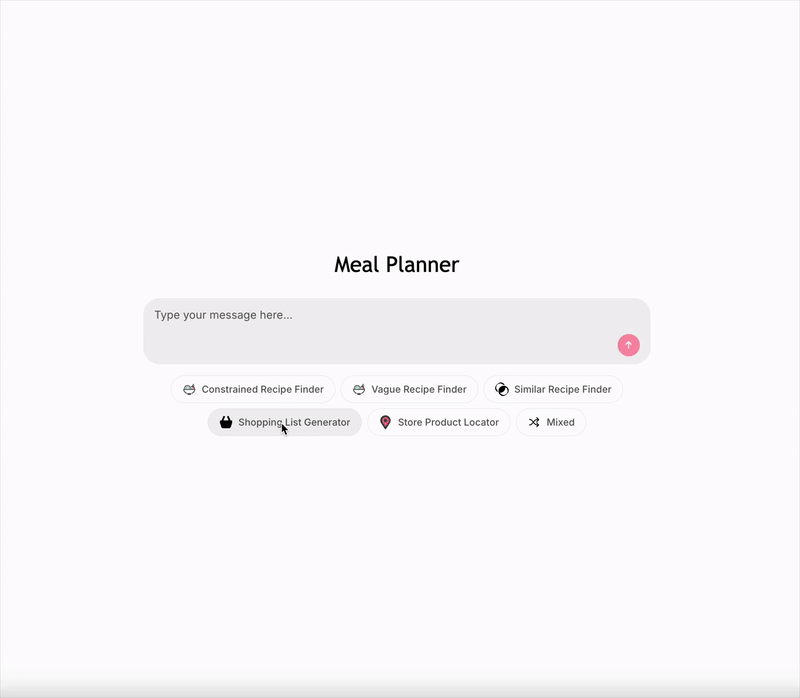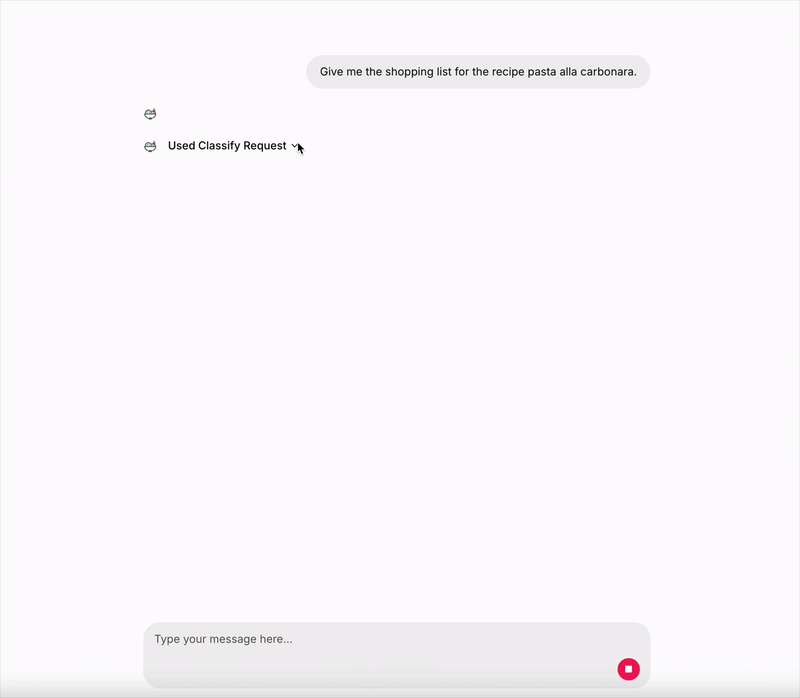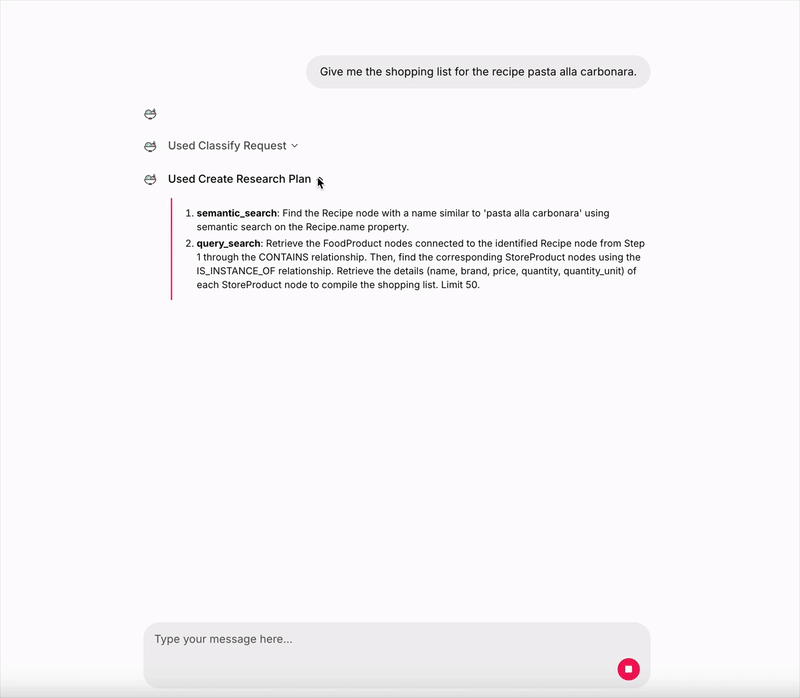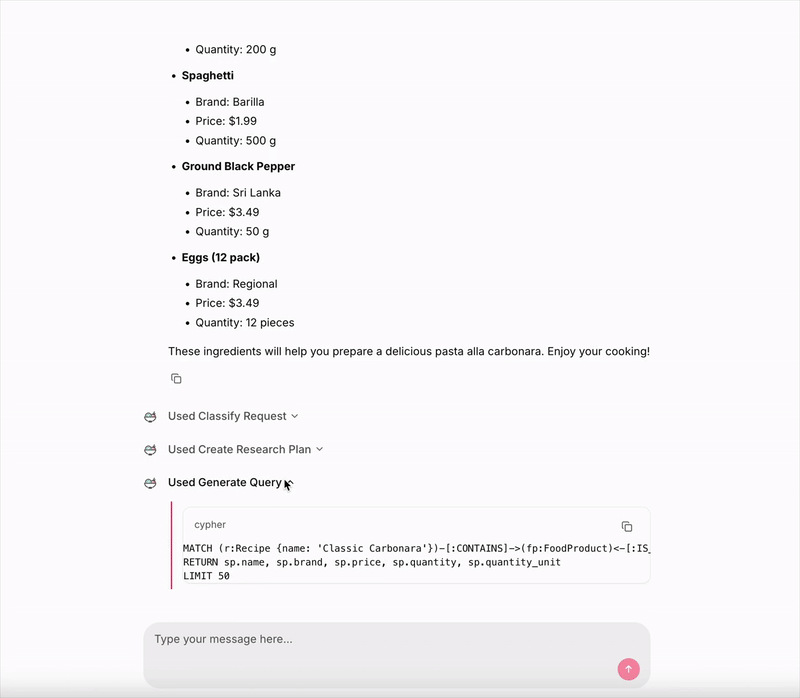
Image by author: Live demo — UI made with Chainlit
Checking from the graph content we see that the complete result is correct.
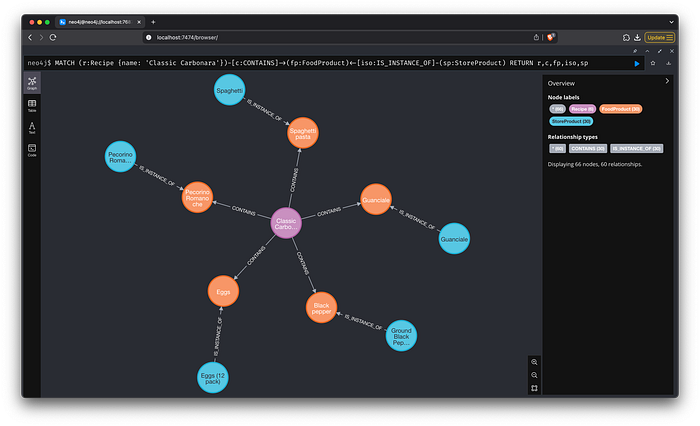
Image by author: Graph response — Store products for “Classic Carbonara”
4. Conclusion
Graph RAG: Technical Challenges and Considerations
Despite the improved performance, implementing Graph RAG is not without its challenges:
- Latency: The increased complexity of agentic interactions often leads to longer response times. Striking a balance between speed and accuracy is a critical challenge.
- Evaluation and Observability: As Agentic RAG systems become more complex, continuous evaluation and observability become necessary.
In conclusion, Graph RAG marks a major breakthrough in the realm of AI. By merging the capabilities of large language models with autonomous reasoning and information retrieval, Graph RAG introduce a new standard of intelligence and flexibility. As AI continues to evolve, Graph RAG will play an fundamental role in various industries, transforming the way we use technology.
References:https://python.langchain.com/docs/how_to/graph_constructing/https://infohub.delltechnologies.com/it-it/p/the-rise-of-agentic-rag-systems/https://langchain-ai.github.io/langgraph/how-tos/https://neo4j.com/labs/genai-ecosystem/llm-graph-builder/https://github.com/Chainlit/chainlit
POSTS ACROSS THE NETWORK

The Rise of AI in Web3 UX: How AI Is Changing Product Design in 2026

10 AI Tools Content Writers Are Quietly Using in 2026 (Beyond ChatGPT)
How AI Agents Are Changing the Way Developers and Creators Approach Video Production

Did AI Write This?
